Deciphering Tumour Microenvironement
A pathologic-genomic approach
Pathological analysis of tumour architecture and cell morphology dates back to the 19th century. Now, morden image-analysis tools promises objective and quantitative insights into hundreds of tumours.
By drawing on the rich information from pathological images and combining it with molecular data from next generation sequencing, we hope to offer more powerful approaches to study the microenvironment and its implications to cancer progression.
Building an automated classifier for histopathological images, enables us to identify heterogeneous cell populations in patient tumours and quantify their spatial proximity to each other.
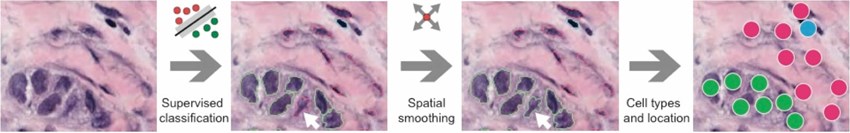
Click to enlarge image
Quantitative scores of cell spatial patterns
The cell spatial patterns can be summarised using methods from spatial statistics and pattern recognition techniques to provide quantitative scores.
These quantitative scores of cell patterns can reveal various metrics that would be impossible to conclude by eye, for example, the degree of clustering and the spatial interactions between different cells.
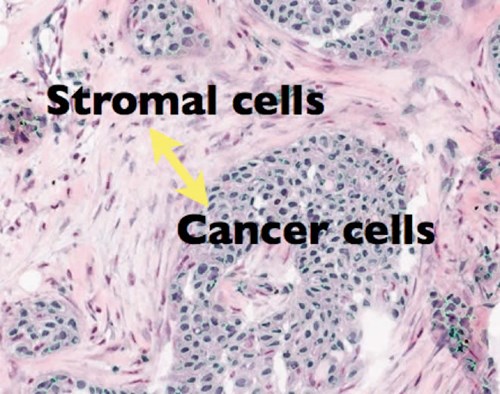
Click to enlarge image
Discovery of functional roles of normal cells
One of the central dogma in cancer biology is the functional roles of heterogenous types of cells in cancer progression.
With quantitative readout from the images, we can correlate the abundance and spatial patterns of cells with patient outcome. For example, by summarising the abundance of lymphocytes in the ER-negative breast cancer, we are able to stratify patients into different outcome groups, where the patients with high numbers of lymphoctyes have significantly better prognosis than patients with few lymphocytes.
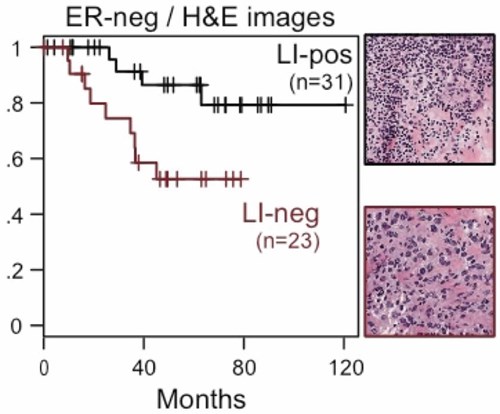
Click to enlarge image
Integrating images with molecular signal
Combining molecular signal from the same samples, the image-based quantitative scores can either complement molecular data, or offer completely new observations from this entirely different angle.
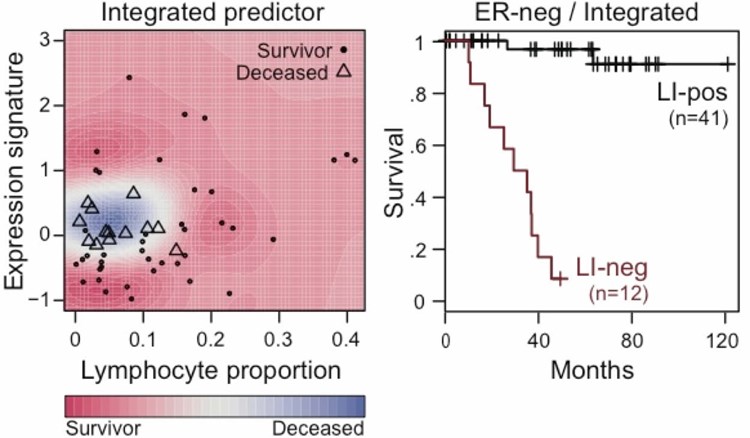
Click to enlarge image